Table of Contents
Assorted evolutionary designs in serious infections
We get started by defining requirements for a persistent an infection. In scientific configurations, a chronic an infection is usually described as a single with both equally prolonged shedding of viral RNA and evidence of infectious virus, possibly through virus isolation in tissue lifestyle or by means of detection of subgenomic RNA. Nevertheless, when surveying numerous reports reporting continual infection, we observed a lack of standardization, with various scientific tests defining chronic infections somewhat inconsistently. Therefore, we expanded our aim to include things like people displaying large-viral-load (VL) shedding for 20 or additional times whilst mining the literature for all this kind of scenarios that ended up accompanied by longitudinal complete-genome sequencing of the virus (Techniques). The criterion of 20 days was primarily based on a meta-assessment of the length of viral shedding (outlined as a beneficial nasopharyngeal polymerase chain response (PCR) take a look at) throughout hundreds of patients identified right up until June 2020, which exposed that suggest duration of upper respiratory tract shedding was around 17 days, with a 95% self-confidence interval ranging from 15.5 days to 18 days16. Of notice, shedding of replication-proficient virus lasted markedly fewer than 20 days. Moreover, estimates of viral shedding are distinctive in some of the more recently detected SARS-CoV-2 variants, this sort of as Delta and Omicron17,18, still, as described under, our evaluation targeted on variants that ended up discovered in previously stages of the pandemic.
Our research yielded a overall of 21 scenario reviews, all of which described people who were diagnosed during 2020 or early 2021, and all of which claimed people who have been infected with viruses belonging to lineages that pre-dated the Alpha variant (Supplementary Table 2). In addition, six patients adhering to the earlier mentioned conditions were being discovered in TASMC, and all out there samples have been sequenced (Procedures). 5 TASMC clients endured from hematologic cancers. The sixth patient suffered from an autoimmune problem and was taken care of with a significant dose of steroids. The 6 TASMC individuals have been all identified in late 2020 or early 2021, with four clients contaminated with a virus from pre-Alpha lineages and two clients infected with a virus from the Alpha lineage (Supplementary Table 2).
Of the 27 chronically contaminated clients (indicate age (s.d.) 55 (21.3) a long time 17/27 male), we inferred that all were being immunocompromised because of to a single or extra of the pursuing: hematologic most cancers (that inherently tends to guide to immunosuppression), immediate anti-B cell cure, substantial-dosage steroid treatment method or really lower CD4+ T cell counts (thanks to AIDS). We noticed really different evolutionary outcomes throughout the array of individuals examined, from appreciable evolution and antibody evasion noticed in some individuals to reasonably static evolution in other folks (Table 1 and Supplementary Tables 1 and 2).
Evolution in chronic bacterial infections as opposed to world-wide transmission chains
We searched for patterns of evolution across all 27 people with chronic infection and as opposed this sample to the pattern noticed beneath (1) generally neutral evolution, in the initial roughly 9 months of viral circulation19,20 (data have been acquired from a sample of ~3,500 sequences created by NextStrain https://nextstrain.org/21 (Methods)) and less than (2) presumed optimistic collection, which transpired in the lineages primary to the 5 at this time defined VOCs (Alpha, Beta, Gamma, Delta and Omicron) (info on lineage-defining mutations (LDMs) of VOCs were obtained from https://covariants.org (Fig. 1a and Supplementary Desk 4)). In each individual scenario, we searched for bins—that is, consecutive locations of 500 bases—enriched for mutations (P < 0.05, binomial test, after correction for multiple testing Methods).
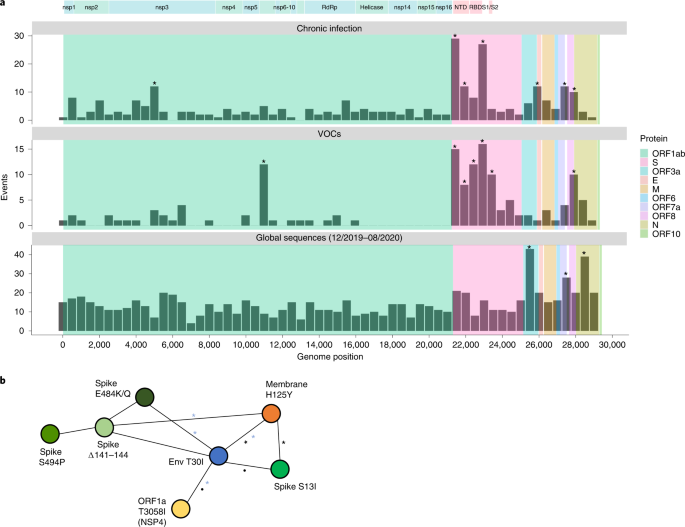
a, Comparison of substitutions observed in chronic infections to VOC LDMs and to substitutions dominated by genetic drift during globally dispersed acute infections. Shown are the number of substitutions observed along the SARS-CoV-2 genome, in bins of 500 nucleotides. The upper panel displays substitutions observed at any timepoint of the 27 chronic infections. The middle panel displays LDMs of the five currently recognized VOCs. The lower panel displays substitutions observed globally during the first 9 months of the pandemic, mostly before the emergence of VOCs. Asterisks mark bins enriched for more substitutions using a one-tailed binominal test, after correction for multiple testing (P < 0.05 Methods and Supplementary Table 8). The genomic positions are based on the Wuhan-Hu-1 reference genome (GenBank ID NC_045512), and the banner on the top shows a breakdown of ORF1a/b into individual proteins and domains of the S protein (see main text). b, A network of co-occurring substitutions across patients with chronic SARS-CoV-2 infection. Each colored circle represents a locus, and a black asterisk and dot represent a significant enrichment under a one-tailed Fisher’s exact test with P < 0.05 and P < 0.1, respectively, after correction for multiple testing. Blue asterisks represent enrichment of co-occurring substitutions in globally observed sequences using a one-tailed X2 test, with P < 0.05 and P < 0.1, respectively, after correction for multiple testing (Methods).
During the first 9 months of virus circulation, we noted that 61% of substitutions were non-synonymous, which is generally what we could expect under lack of both positive and purifying selection and in line with reports suggesting incomplete purifying selection during the early stages of SARS-CoV-2 spread22. During this time, we observed a relatively uniform distribution of substitutions across most of the genome, with some enrichment in ORF3a, ORF7a, ORF8 and N. This enrichment was previously reported and may be due to more relaxed purifying selection in these regions or higher mutation rates19 adaptive evolution at these regions also cannot be ruled out.
In general, the patterns obtained in chronic infections and in the LDMs of VOCs were very similar. The average proportion of non-synonymous substitutions in chronic infections and LDMs of VOCs was 78% and 82%, respectively, which was much higher than that observed during the first stage of the pandemic and generally suggestive of positive selection. On the other hand, we see less similarity between mutations in chronic infections and mutations that fix after a VOC has emerged (Supplementary Fig. 1), with a much lower proportion of non-synonymous substitutions in the latter (on average, 61%). A likely explanation for this observation is that after a VOC spreads in the population, selection is more limited due to the very tight transmission bottleneck9,10,11,12.
The most striking similarity between chronic infections and VOC LDMs was observed along the S protein and, in particular, at the regions that correspond to the N-terminal domain (NTD) (genomic nucleotides 21,598–22,472) and the receptor-binding domain (RBD) (genomic nucleotides 22,517–23,183). Several mutations at the RBD have been shown to enhance affinity to the ACE2 receptor and allow for better replication23,24, whereas other mutations, both at RBD and NTD, are known to enhance antibody evasion25,26,27. The most commonly observed substitutions in chronic infections were in the S protein: E484K/Q and various deletions in the region spanning the NTD supersite, particularly amino acids 140–145, all shown previously to confer antibody evasion28. Chronic infections shared the enrichment of ORF3a/ORF7a/ORF8 mutations with the ‘neutral’ set but lacked an enrichment across most of the N protein. Overall, it seems that mutations in chronic infections are predictive of LDMs of VOCs, as was noted previously2.
When focusing on the differences between VOCs and viruses in chronic infections, several intriguing differences emerged. First, four VOCs bear a three-amino-acid deletion in the nsp6 protein (ORF1a:∆3,675–3,677), which is an event not observed in our set of chronic infections. Next, in VOCs, there is an enrichment in the region of the S encompassing the S1/S2 boundary (positions 23,500–24,000 in Fig. 1a). This enrichment is primarily driven by S:P681H/R, a highly recurrent globally occurring mutation29, surprisingly never observed in our chronic infection set. A recent study analyzed recurrent mutations, with recurrence indicative of positive selection, and tested which of the recurrent mutations led to clade expansion—that is, were associated with onwards transmission30. Some recurrent mutations led to more dense clades, suggesting that they were especially successful in driving transmission, whereas others did not lead to considerable onwards transmission, suggesting that they were less successful. Notably, we observed that successful recurrent mutations were almost never present in our chronic set, whereas less successful recurrent mutations (S:E484K/Q and S:∆144) were the most abundant (Table 2). Overall, these results suggest that there may be a tradeoff between antibody evasion and transmissibility. This tradeoff, if it exists, might not play a role in chronic infections but would affect the ability of a variant created in a chronic infection to be transmitted onwards. Thus, only under specific conditions, a transmissible variant would emerge in chronic infections. Four of five VOCs independently acquired a mutation at or near the S1/S2 boundary (S:P681H/R or H655Y), suggesting that this may be a factor driving transmissibility. We note that Beta is an exception with no such mutations, yet this variant also displayed limited global transmission.
We went on to examine co-occurring substitutions, defined as pairs of substitutions that appeared in two or more patients. We used Fisher’s exact test to assess whether pairs of substitutions occurred together more often than expected from their individual frequencies (Methods) as a measure of possible epistasis. Intriguingly, four pairs of substitutions across four different proteins emerged as significantly enriched and formed a network of interactions: T30I in envelope, H125Y in the membrane glycoprotein, S13I in the S protein and T3058I in ORF1a (Fig. 1b). This finding was intriguing on multiple fronts. First, envelope and membrane glycoprotein have generally remained very conserved throughout the entire pandemic, and, specifically, the two replacements found are at highly conserved sites (Supplementary Table 1). However, despite their rarity, we found that some of the pairs of mutations also tend to significantly co-occur in globally dispersed sequences (blue asterisks in Fig. 1b). The replacements in S and ORF1a, on the other hand, have been observed only a small number of times in the global phylogeny. Notably, all of the first three proteins form a part in the virion structure itself however, the functional meaning of this remains unclear. Other pairs of mutations found to co-occur were the three most common S antibody evasion mutations, yet these co-occurrences were not statistically significant. Larger cohorts of patients and further data will be required to determine the implications of these findings.
Correlates of antibody evasion
We noted very wide variation in the background and treatments given to different patients, both for their background condition and for Coronavirus Disease 2019 (COVID-19). When examining medical background, the patients could be roughly classified into one of the following categories: hematologic cancers, HIV/AIDS, organ transplantation and autoimmune disorders (Table 1). The latter two categories were often treated with steroids. Some, but not all, of the patients with hematological cancer and others were treated with antibodies targeting B cells, presumably causing profound B cell depletion. In line with this, most of the patients with confirmed B cell depletion showed negative serology for SARS-CoV-2 at one or more timepoints (Supplementary Table 1). Some patients were treated with ABT against SARS-CoV-2, whereas others were not and, in some ABT-treated patients, antibody evasion mutations were detected, whereas, in others, they were not. Finally, we found that, whereas in some ABT-treated patients, antibody evasion mutations were detected, sometimes these mutations fixed before the treatment. The course of VL across time, coupled with ABT, is illustrated for some patients in Fig. 2b. Thus, for example, patient P5 and the patient described by Choi et al.8 are shown to fix antibody evasion mutations just before ABT.
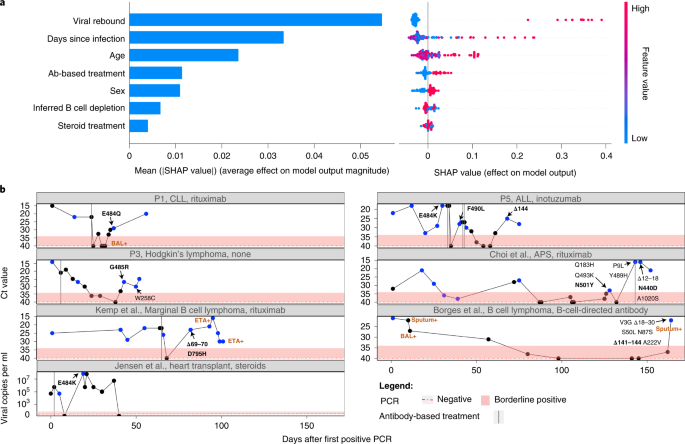
a, Results of a random forest classifier used to explain an outcome of antibody evasion. The effect of each feature on model outcome is shown: mean SHAP absolute values (left) and individual SHAP values for each feature, ordered based on contribution (right). The color range corresponds to the values of each feature, from red (high value) to blue (low value). b, Illustration of individuals who experienced viral rebound and mutations associated with antibody evasion. Ct values are used here as an inversed proxy for VL and are presented according to the day of infection (denoted as number of days after the first positive PCR test), with the dashed red horizontal line and shaded area representing a negative or borderline result, respectively. Blue dots represent samples that were sequenced. Only amino acid replacements in the S protein are shown, with predicted antibody evasion mutations shown in bold (Supplementary Table 1). Positive samples from BAL, ETA or sputum are indicated in brown. Antibody-based anti-COVID-19 treatments are represented by dashed vertical lines on the day of administration. ALL, acute lymphoblastic leukemia APS, antiphospholipid syndrome CLL, chronic lymphocytic leukemia ETA, endotracheal aspirates P, patient.
We noted that many patients (four of the six patients sequenced herein and several others in the total set of 27 patients) displayed an intriguing cycling pattern of VL (reflected by cycle threshold (Ct) values), with very high Ct values reaching negative or borderline-negative results at one or more stages of the infection, followed by rebound of the virus (Fig. 2b). In the four above-mentioned patients, this rebound was accompanied by clinical evidence of disease, which is highly suggestive of active viral replication. Several different hypotheses could explain this pattern. First, the virus may have cleared and been followed by re-infection with another variant. Because this pattern can be ruled out using sequencing, such cases were excluded from our analysis (Methods). Second, the virus may cycle between different niches, such as upper and lower airways. Its re-emergence in the upper airways (nasopharynx) may be due to selective forces or genetic drift. When considering selective forces, viral rebound may occur due to the near clearance of the virus, driven either by ABT or by the endogenous immune system, and followed by the emergence of a more fit variant with antibody evasion properties.
We fit a random forest classifier to assess the effect of different clinical and demographic features on an outcome of antibody evasion (Methods and Supplementary Tables 2 and 3). We treated each sequencing timepoint as a sample and used age, sex, B cell depletion, steroid treatment, days-since-infection, ABT and viral rebound as explaining variables. We then trained a classifier while considering the structure of the data, composed of samples belonging to the same patient (Methods). After training, we generated SHapley Additive exPlanations (SHAP) values31,32 that quantified the effect of each feature on the classifier’s outcome. We found that the feature with the strongest association with antibody evasion was viral rebound, followed by days-since-infection and age (Fig. 2a). Other features had a relatively minor effect, and similar results were obtained with other classifiers (Supplementary Figs. 2 and 3). Regarding the effect of age, we note that young individuals are a minority in this dataset and rarely present an antibody evasion mutation, and, thus, the small sample size may be responsible for the small effect observed with this feature. All in all, these results suggest that ABT is not necessary for driving antibody evasion, in line with the fact that evasion is sometimes observed before (for example, E484K in P5 Fig. 2b) or in the absence of ABT (for example, ref. 33). If so, what may be driving immune escape in some patients is actually the weakened immune system of the patient, although ABT and its waning may also play a role in some patients. To summarize, viral rebound may serve as an indicator for the emergence of a mutant with properties of antibody evasion (Fig. 2b), and monitoring for viral rebound in patients with chronic disease is critical.
Next, we went on to examine patterns of variation over time across the different patients. In many of the case reports, the authors noted the emergence and disappearance (and sometimes re-emergence) of particular substitutions (Fig. 3). For example, in patient B reported by Perez-Lago et al.34, the mutation S:A1078V is present at a low frequency on day 81, rises to fixation on day 100 and then drops and disappears from day 107 onwards (Fig. 3). When re-analyzing the data, we noted that this pattern of dynamic polymorphisms across time was observed in most patients (Supplementary Table 2). From an evolutionary point of view, it is quite unlikely for one or more substitutions to disappear from a given population, and, because we observe this at very different loci across all patients, we consider that it is not likely that all of this pattern is due to recurrent sequencing problems or due to biases of the viral polymerase. We and others have previously noted sequencing errors that occur predominantly when VL is low, when errors that occur during reverse transcription or early PCR cycles are carried over to higher frequencies10,11,35. However, this phenomenon most often leads to errors in intra-host variants segregating at relatively low frequency and is less common at the consensus sequence level, which is defined here as mutations present at a frequency of 80% or higher. We, thus, conclude that the existence of dynamic polymorphisms likely reflects subpopulations of the virus that co-exist in a patient’s body, as further discussed below.
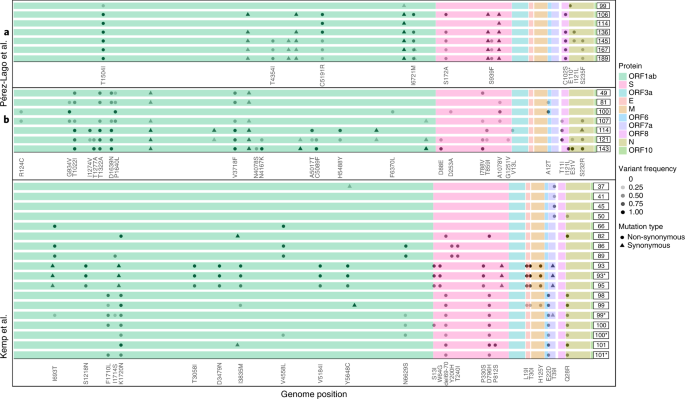
Each series of boxed lines represents a patient, and each line represents a sequenced timepoint with time-since-infection on the right. The different open reading frames are color-coded. For each patient, only mutations relative to the first timepoint sequenced that appeared at a frequency ranging from 20% to 100% are shown. Most samples were nasopharyngeal, except those marked by asterisks, which were obtained from endotracheal aspirates.





More Stories
Top Natural Pain Management Techniques Used by Florida Doctors That Actually Work
Most medical college students get worried abortion rules will ‘hinder their foreseeable future care’
Edinburgh to host supercomputer process that may perhaps advance drugs, AI and strength